AICult: Artificial Intelligence with Cultured Neural Networks
Aim of the AICult project is to investigate and develop solutions to use Cultured Neuronal Networks (CultNN), that is biological cultures of real neuronal cells, both normal or carrying mutations associated with cognitive diseases, for Artificial Intelligence (AI). We will investigate training methods for CultNN, learning optimized solutions for engineering and interacting with CultNN. We will leverage existing state of the art technology for combining calcium imaging with light stimulation (Optogenetics), to connect computers to CultNN.
We propose a radical paradigm shift for AI, based on the direct use of CultNN, rather than bioinspired Artificial Neural Networks (ANN). We aim at designing devices based on real biological cultures of neuronal cells capable of learning and performing AI tasks. The final product of our research will be a “biological” AI co-processor that can perform AI tasks and comparisons between normal CultNN and CultNN of neurons carrying genetic mutations associated with cognitive diseases.
To achieve our aim we will integrate competences and do research along the following multidisciplinary lines:
Artificial Intelligence:
- digital models of CultNN, to be able to perform simulated experiments and predict expected performance of CultNN.
- AI training protocols dedicated to CultNN.
Neural Computation and Neuroscience:
- signal processing analysis, to infer complex network interactions in CultNN.
- analysis of electrical activity at: 1) single-cell level, to infer basic signal processing of biological neurons in a CultNN; 2) network level, by imaging of calcium changes in response to optogenetically-induced voltage changes.
Cell biology:
- CultNN engineering with:
- 1) light-gated ion channels to control neuronal electrical activity by optogenetics;
- 2) genetically encoded calcium indicator gcAMP to record neuronal activity.
- Development of protocols of neuronal differentiation to produce cultures with a desired composition, density and physiological maturity degree of both inhibitory and excitatory neurons.
AICult will open up new scientific opportunities and have an impact in the long term:
- Surpassing the scale limitation of ANN, which perform just a small fraction of the tasks executed by a Human Brain, still taking much more space and resources.
- Reducing energy and pollution footprint: the human brain, so CultNN, consume orders of magnitude less energy (around 20W on average) than ANN (Strubell E. et alt. 2019), which produce high carbon emission and pollution
- Gaining additional understanding of the biological brain.
- Creating trained CultNN that can act as both retina and visual cortex, leveraging on their optogenetics properties.
- In vitro modeling of cognitive diseases, by comparing the AI ability of normal CultNN to those made of neurons carrying disease-associated genetic mutations
AICult relies on a multidisciplinar consortium with very complementary competences. The consortium covers the following disciplines: Computer Science, Artificial Intelligence, Neural Computation, Neurosciences, Cell Biology, Molecular Biology, and Electrophysiology, which will be integrated as depicted in the following figure:

Our research methodology is depicted in the figure below.
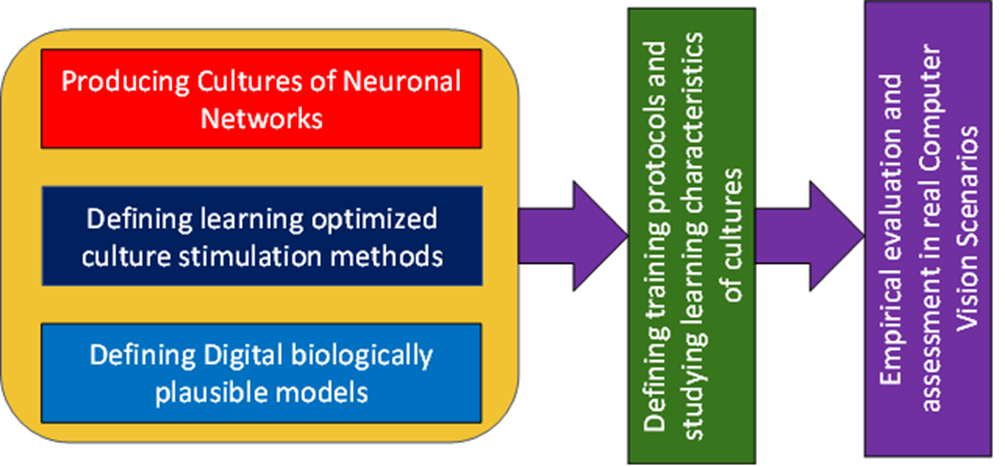
We will advance knowledge along three parallel, yet interacting, research lines:
- producing learning optimized cultures to execute AI tasks
- defining learning optimized culture stimulation methods
- defining digital biologically-plausible models of cultures to simulate learning processes.
Results of these three research lines will allow
- Advancing knowledge in the study of training protocols and learning paradigms on CultNNs
- Performing incrementally difficult experiments on Computer Vision tasks.
Pubblications
| # | Partner | Publication |
|---|---|---|
| 1 | CNR, SNS | L. Iannello, F. Tonelli, L. M. Calcagnile, F. Cremisi, R. Mannella and A. D. Garbo, “Analysis of MEA recordings in cultured neural networks,” 2024 IEEE Workshop on Complexity in Engineering (COMPENG), Florence, Italy, 2024, pp. 1-5, doi: 10.1109/COMPENG60905.2024.10741515. |
| 2 | CNR, SNS | Ludovico Iannello, Luca Ciampi, Gabriele Lagani, Fabrizio Tonelli, Eleonora Crocco, Lucio Maria Calcagnile, Angelo Di Garbo, Federico Cremisi, Giuseppe Amato, From Neurons to Computation: Biological Reservoir Computing for Pattern Recognition, 32nd International Conference on Neural Information Processing (ICONIP) 2025, November 20-24, 2025, Okinawa Institute of Science and Technology (OIST), Japan, also arXiv:2505.03510v2, 2025 |
| 3 | CNR, SNS | Tonelli, Fabrizio, Iannello, Ludovico, Gustincich, Stefano, Di Garbo, Angelo, Pandolfini, Luca, Cremisi, Federico, Dual inhibition of MAPK/ERK and BMP signaling induces entorhinal-like identity in mouse ESC-derived pallial progenitors, Stem Cell Reports, Elsevier, Volume 20, Issue 2, 2025/02/11, doi: 10.1016/j.stemcr.2024.12.002 |
| 4 | CNR | Lagani, G., Falchi, F., Gennaro, C., Amato, G. (2025). CA3D: Convolutional-Attentional 3D Nets for Efficient Video Activity Recognition on the Edge. In: Del Bue, A., Canton, C., Pont-Tuset, J., Tommasi, T. (eds) Computer Vision – ECCV 2024 Workshops. ECCV 2024. Lecture Notes in Computer Science, vol 15633. Springer, Cham. https://doi.org/10.1007/978-3-031-91979-4_18 |
| 5 | CNR, SNS | Eleonora Crocco, Ludovico Iannello, Fabrizio Tonelli, Gabriele Lagani, Luca Pandolfini, Giuseppe Amato, Angelo Di Garbo, Federico Cremisi, A proper Excitatory/Inhibitory ratio is required to develop synchronized network activity in mouse cortical cultures, bioRxiv 2025.02.28.640720; doi: https://doi.org/10.1101/2025.02.28.640720 |
| 6 | SISSA | Antonello Mallamaci, Osvaldo Artimagnella, Gabriele Liuzzi, Foxg1 and companions: not only transcription factors, bioRxiv 2024.12.16.628377; doi: https://doi.org/10.1101/2024.12.16.628377 |
| 7 | CNR | Gabriele Lagani, Fabrizio Falchi, Claudio Gennaro, Hannes Fassold, Giuseppe Amato, Scalable bio-inspired training of Deep Neural Networks with FastHebb, Neurocomputing, Volume 595, 2024, 127867, ISSN 0925-2312, https://doi.org/10.1016/j.neucom.2024.127867. |
| 8 | CNR,SNS | Ludovico Iannello, Fabrizio Tonelli, Federico Cremisi, Lucio Maria Calcagnile, Riccardo Mannella, Giuseppe Amato, Angelo Di Garbo, Criticality in neural cultures: Insights into memory and connectivity in entorhinal-hippocampal networks, Chaos, Solitons & Fractals, Volume 194, 2025, 116184, ISSN 0960-0779, https://doi.org/10.1016/j.chaos.2025.116184. |
| 9 | CNR,SNS | Ludovico Iannello, Luca Ciampi, Fabrizio Tonelli, Gabriele Lagani, Lucio Maria Calcagnile, Federico Cremisi, Angelo Di Garbo, Giuseppe Amato, From Neural Activity to Computation: Biological Reservoirs for Pattern Recognition in Digit Classification, 2nd Human-inspired Computer Vision ICCV 2025 Workshop, 20th October 2025, Honolulu, Hawaii |
| 10 | SISSA | Osvaldo Artimagnella, Elena Sabina Maftei, Maurizio Esposito, Remo Sanges, Antonello Mallamaci A (2024). Foxg1 regulates translation of neocortical neuronal genes, including the main NMDA receptor subunit gene, Grin1. BMC Biol. 22(1), 180. doi: 10.1186/s12915-024-01979-x. |
| 11 | SISSA | Maria Ayub and Antonello Mallamaci. Foxg1 promotes neuron allocation to engrams and facilitates fear memorization. bioRxiv 2026. https://doi.org/10.64898/2026.02.13.705789 |
| 12 | CNR,SNS | Luca Ciampi, Ludovico Iannello, Fabrizio Tonelli, Gabriele Lagani, Angelo Di Garbo, Federico Cremisi, Giuseppe Amato, Neuro-Inspired Visual Pattern Recognition via Biological Reservoir Computing, 2026, https://doi.org/10.48550/arXiv.2602.05737 |
Partners
- The Consiglio Nazionale delle Ricerche (CNR) research unit will participate in the project with Institute of Information Science and Technologies (CNR-ISTI) collaborating with the Institute of Neurosciences (CNR-IN).
- PI: Giuseppe Amato
- Scuola Normale Superiore (SNS), Laboratory of Biology
- PI: Federico Cremisi
- Scuola Internazionale Superiore di Studi Avanzati (SISSA), Laboratory of Cerebral Cortex Developement
